
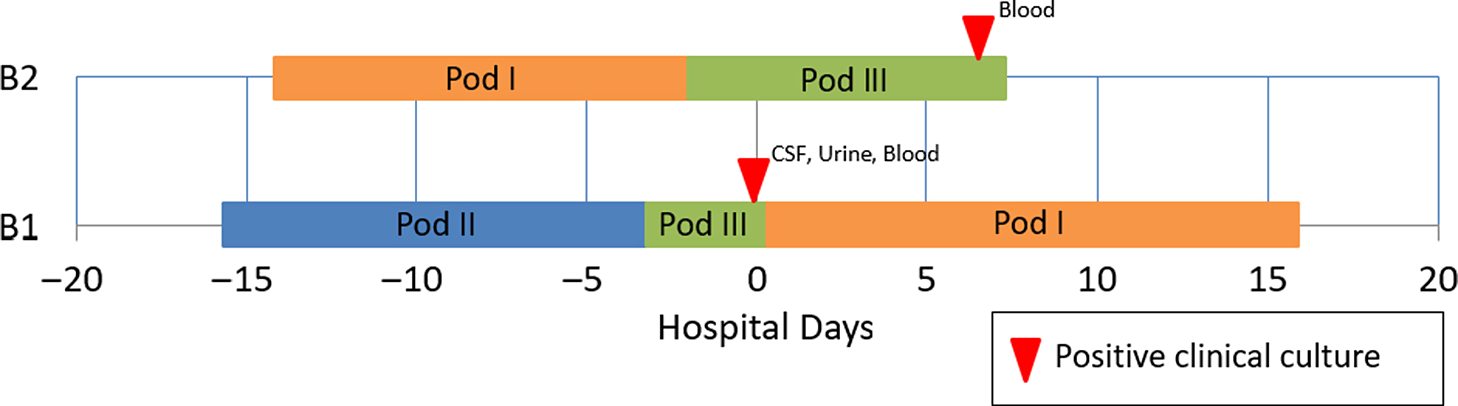
Whole-genome sequencing for neonatal intensive care unit outbreak investigations: Insights and lessons learned, Antimicrobial Stewardship & Healthcare Epidemiology

New ERGO Feature: Amplicon Sequence Analysis Workflow — Igenbio

Metabolism of ʟ -arabinose converges with virulence regulation to promote enteric pathogen fitness

PDF) SPANDx: A genomics pipeline for comparative analysis of large haploid whole genome re-sequencing datasets

Reduced Representation Methods for Subgenomic Enrichment and Next-Generation Sequencing

Investigation of Respiratory Syncytial Virus Outbreak on an Adult Stem Cell Transplant Unit by Use of Whole-Genome Sequencing

Number of SNVs called by GATK-UGT with different cutoffs of genotype

Figure 2 from The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data.
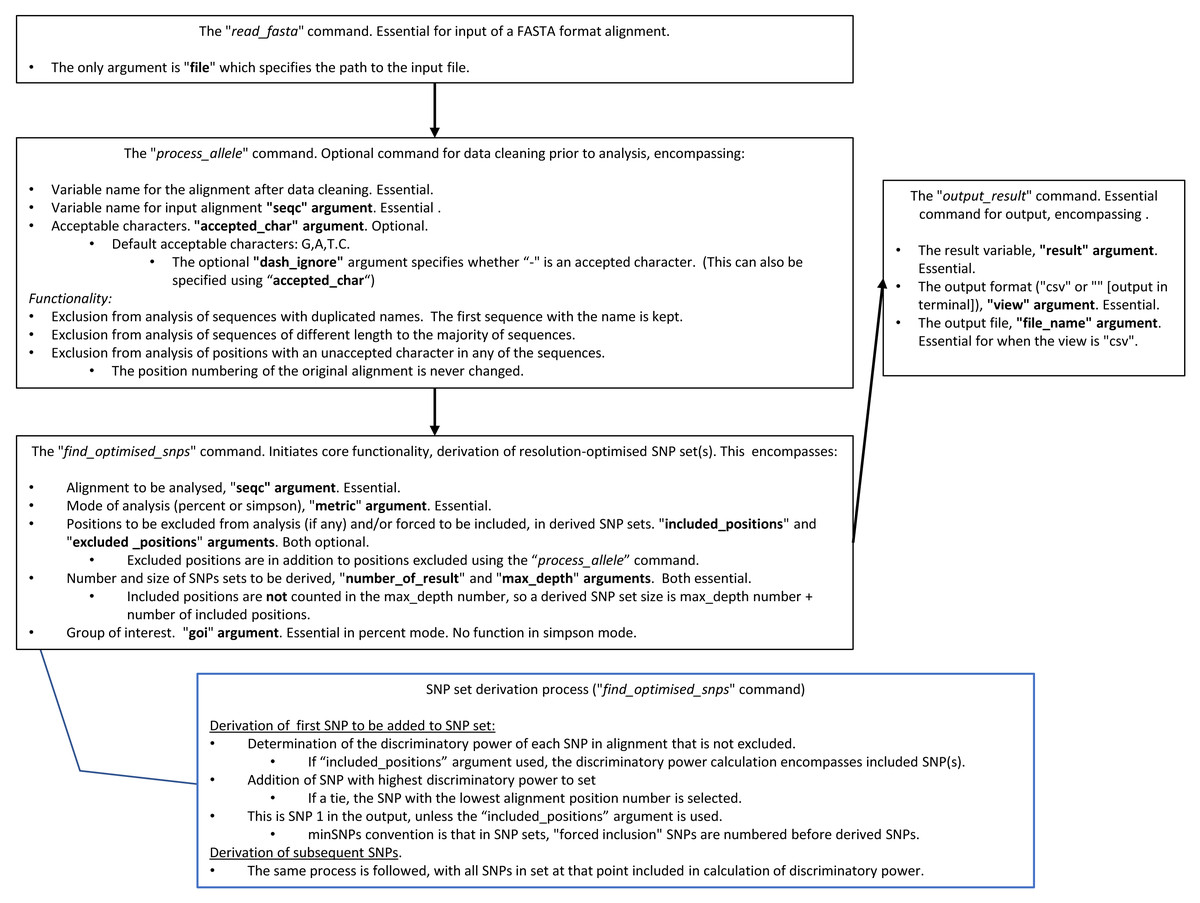
minSNPs: an R package for the derivation of resolution-optimised SNP sets from microbial genomic data [PeerJ]

SPANDx workflow for analysis of haploid next-generation re-sequencing data.

SPANDx workflow for analysis of haploid next-generation re-sequencing data.